CRISPR
CRISPR (clustered regularly interspaced short palindromic repeats) sequences are clusters of DNA that recognize invading cells and deploy an enzyme to cut apart the RNA nucleotides of the invaders. They’re a basic immune system device, detecting and cleaving apart viruses in prokaryotic organisms.
Researchers use CRISPRs for genetic modification. They are uniquely valuable to geneticists because of their two stages: first, finding and targeting specific strands of DNA and then, second, deleting or replacing that DNA through reverse transcription. These two stages of CRISPR sequences are far more efficient and accurate than previous methodologies. Now researchers can mutate multiple genes at once, a process that previously took months or years, which is necessary for research into single illnesses caused by multiple genetic defects. Already, researchers have used CRISPRs to modify agricultural plants, making them more nutritious, pestilence-resistant, and drought-tolerant. In mice, CRISPRs have been used to correct genetic errors including those responsible for sickle-cell anemia, muscular dystrophy, and an error associated with cystic fibrosis.
CRISPRs were first used to modify human cells in 2012, prompting rapid technological transformations in the field of genetics and raising ethical questions about engineering human genes.
Contents
Parts of CRISPR Sequences
CRISPR sequences in microbial hosts are composed of three parts, CRISPR-associated (Cas) genes that are the cutting enzymes, tracrRNA, and CRISPR RNA (crRNA). Together, tracrRNA and crRNA are known as guide RNA (gRNA). Some CRISPR complexes also include a strand of repair DNA to be inserted into the target.
Targeting
The gRNA strand handles targeting in the CRISPR complex. gRNA is approximately 80 base pairs and binds to the target genome to direct the cutting enzyme. Three types (I-III) of CRISPR mechanisms have been identified, but type II is the most commonly used and is described here.
 In microbes, the tracrRNA binds to the DNA of invading phages and plasmids, essentially invading viruses. In studies, researchers customize the tracrRNA to find and target whichever genes they want to edit, encoding the 20 base pairs of tracrRNA to match whatever target complementary sequence they want to bind to in the target genome.
In microbes, the tracrRNA binds to the DNA of invading phages and plasmids, essentially invading viruses. In studies, researchers customize the tracrRNA to find and target whichever genes they want to edit, encoding the 20 base pairs of tracrRNA to match whatever target complementary sequence they want to bind to in the target genome.
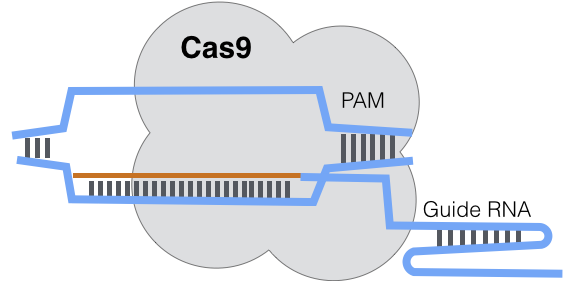
The target sequence must be immediately upstream of a Protospacer adjacent motif (PAM) strand of DNA in the target genome. A PAM in the target genome is essentially a flag and a bookend, flagging the bacteria as a target and stopping the binding when the gRNA reads the PAM.
Once the Cas-gRNA complex (the CRISPR sequence) is expressed, it denatures the target DNA and anneals to it. This is referred to as a double-strand break (DSB). Essentially, the Cas-gRNA complex splits the two strands of DNA apart and binds to each strand. If the tracrRNA and the target DNA are sufficiently similar, the Cas enzyme cuts the target DNA.
Cutting Enzymes
Cas are nuclease enzymes, the cutting enzymes in CRISPRs. The most commonly used variant of Cas in research settings is Cas9. It is derived from a species of streptococcal bacteria. Cas9 cuts the target DNA through the combined activity of two endonuclease domains, RuvC and HNH.
The goal of the CRISPR system is to mutate the target strand in some way, either to make it non-functioning or to repair it in some way. So once the Cas enzyme has cut the target, it inserts new DNA via the optional strand of repair DNA, deletes existing genes, or frameshifts (moving existing base pairs to other places on the strand).
Next Cas rejoins the split strands of DNA using either the non-homologous end joining (NHEJ) pathway or the homology directed repair (HDR) pathway
Randall Platt successfully implanted the Cas9 enzyme into mice embryo, making it a permanent part of the mouse’s genome. The Cas9 mouse has become an essential tool in CRISPR research, allowing researchers to target multiple DNA strands at once and mutate them all.
In general, CRISPR research is being conducted on mice. Since the 1980s, genetically modified mice have served as the standard test subject. Humans and mice shared a common ancestor approximately 80 million years ago, and today the mouse genome is considered very similar to the human genome, with estimates ranging between 70 percent and 85 percent. Mice also develop complex diseases that affect their immune system and the brain.
Effectiveness
There is potential for error in the CRISPR process at multiple stages. For one, gRNA can go off target, binding to non-identical strands of target DNA, modifying the wrong sequence. In research settings, it’s best for the gRNA to be identical to the target DNA and with no other homologous strands in the target genome. Also, when rejoining the target DNA, the non-homologous end joining (NHEJ) pathway frequently results in small random insertions or deletions.
CRISPR can be used to edit and delete certain specific regions of DNA. The editing/modification by CRISPR of which type of DNA region is most likely to be error-prone?
A. Centromeric sequences
B. Small RNA cluster regions
C. Short tandem repeats
D. GAPDH open reading frame
Question to be considered: Propose some ways to improve the CRISPR process to avoid these errors. Justify your answer.
Previously-used, Genome-editing Technologies
Previously, Zinc fingers (ZFs) and transcription activator-like effectors (TALEs) were the primary tools for gene editing. But both required building customized DNA-binding protein for every genomic target. They were expensive and time-consuming. Whereas, CRISPRs use the standard Cas cutting enzyme (typically Cas9) and a short gRNA sequence that was spliced from the targeted genome. Essentially researchers can attach the mutated gene they want to target to a standard cutting tool, and then set it to delete or mutate the target DNA.